class: top, right, inverse, title-slide # Multiple logistic regression looking at wines ### Zach Mullis, Matt Goard, Ben Tibbits, Sebastien Lee ### updated: 2021-03-15 --- class: inverse, center, middle --- Dr A's comments Model: ```r wines2 <- wines %>% mutate(color = ifelse(color == "red", 1, 0)) # str(wines2) modA <- glm(color ~ sulphates, data = wines2, family = "binomial") summary(modA) ``` ``` Call: glm(formula = color ~ sulphates, family = "binomial", data = wines2) Deviance Residuals: Min 1Q Median 3Q Max -2.7623 -0.6361 -0.4279 -0.2144 2.5790 Coefficients: Estimate Std. Error z value Pr(>|z|) (Intercept) -6.4050 0.1684 -38.03 <2e-16 *** sulphates 9.4425 0.2872 32.88 <2e-16 *** --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 (Dispersion parameter for binomial family taken to be 1) Null deviance: 7251.0 on 6496 degrees of freedom Residual deviance: 5617.5 on 6495 degrees of freedom AIC: 5621.5 Number of Fisher Scoring iterations: 5 ``` --- ```r nullmod <- glm(color ~ 1, family="binomial", data = wines2) PR2 <- 1 - logLik(modA)/logLik(nullmod) PR2 # McFadden's R2 --- Can we get a better model? ``` ``` 'log Lik.' 0.2252801 (df=2) ``` ```r modB <- glm(color ~ density + chlorides, data = wines2, family = "binomial") # summary(modB) PR2 <- 1 - logLik(modB)/logLik(nullmod) PR2 # McFadden's R2 --- Can we get a better model? ``` ``` 'log Lik.' 0.3733582 (df=3) ``` ```r modC <- glm(color ~ ., data = wines2, family = "binomial") # summary(modC) PR2 <- 1 - logLik(modC)/logLik(nullmod) PR2 # McFadden's R2 --- Can we get a better model? ``` ``` 'log Lik.' 0.9414927 (df=13) ``` --- How well are we doing? ```r wines$color <- as.factor(wines$color) WT <- predict(modC, type ="response") WT <- (ifelse(WT > .5, "red", "white")) p_class <- factor(WT, levels = c("red", "white")) library(caret) confusionMatrix(p_class, wines$color) ``` ``` Confusion Matrix and Statistics Reference Prediction red white red 1580 11 white 19 4887 Accuracy : 0.9954 95% CI : (0.9934, 0.9969) No Information Rate : 0.7539 P-Value [Acc > NIR] : <2e-16 Kappa : 0.9875 Mcnemar's Test P-Value : 0.2012 Sensitivity : 0.9881 Specificity : 0.9978 Pos Pred Value : 0.9931 Neg Pred Value : 0.9961 Prevalence : 0.2461 Detection Rate : 0.2432 Detection Prevalence : 0.2449 Balanced Accuracy : 0.9929 'Positive' Class : red ``` --- ```r ggplot(data = wines2, aes(x = sulphates, y = color)) + geom_jitter(width = 0, height = 0.05, alpha = 0.1) + theme_bw() + geom_smooth(method = "glm", se = FALSE, method.args = list(family = binomial(link = "logit")), color = "red") ``` 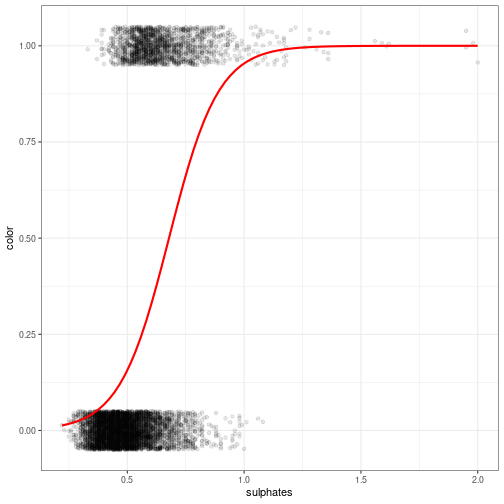<!-- --> --- # assumptions of logistic functions: --- # Here is a plot of the residuals <img src="xariganSlides_files/figure-html/unnamed-chunk-5-1.png" style="display: block; margin: auto;" /> --- class: inverse, center, middle # We discovered that... --- # Using `ggplot2` to illustrate Sulphates, color, and quality together <img src="xariganSlides_files/figure-html/unnamed-chunk-6-1.png" style="display: block; margin: auto;" /> ??? Write everything you want to say about the model here. Pressing shift p in the slides will put you in presentation mode. --- # Showing Quality as a relationship of alcohol between red and whites <img src="xariganSlides_files/figure-html/unnamed-chunk-7-1.png" style="display: block; margin: auto;" /> ## In conclusion: ??? if you want to hide notes that you can use later, doing ??? lets you have a hidden section